 P-values are from a linear model incorporating a binary variable encoding full HLA-I heterozygosity, and age as a continuous variable. e Association of full HLA-I heterozygosity (6 different HLA-I alleles) with number of unique CDR3s in CMV- individuals full HLA-I heterozygosity P = 0.02, estimate = 29,248.4 age P = 0.002, estimate = − 1342.6. P-values are from a linear model incorporating number of unique HLA-I alleles and age. d Association of HLA-I polymorphism with increased number of unique CDR3s in CMV- individuals HLA-I P = 0.02, estimate = 18,787.8 age P = 0.002, estimate − 1326.3. c Association of CMV seropositivity (CMV+) with reduced TCR repertoire diversity (number of unique CDR3s). b Association of CMV seropositivity (CMV+) with reduced TCR repertoire diversity (Shannon entropy). a Variation in number of unique CDR3s and Shannon entropy, two measures of TCR repertoire diversity, across the cohort. Altogether, these data suggest that CMV diminishes TCR repertoire diversity, and are in line with a recent study demonstrating dramatic reduction of the antibody repertoire after measles infection -highlighting the need for widespread and continuous vaccination against infectious disease.ĬMV serostatus and HLA-I genotype are associated with TCR repertoire diversity. 1b), consistent with prior work demonstrating that CMV alters the diversity, but not overall size of the CD8+ T cell response. This reduction was most striking when using Shannon entropy ( P < 0.0001, Wilcoxon test Fig. Indeed, individuals with CMV (CMV+) exhibited a reduction in TCR repertoire diversity compared to those without (CMV-) (Fig. 
Accordingly, we anticipated that CMV-a chronic infection prevalent in 30–90% of adults and a model system for the study of public T cell responses -would be a key determinant of the observed wide variation in repertoire diversity. We found both measures to be highly correlated, and observed high variability in TCR repertoire diversity across the cohort (Fig. richness), and Shannon entropy, a diversity measure that is weighted by the abundance of each CDR3. We first quantified TCR repertoire diversity by applying two measures widely used in repertoire and ecological studies-the number of unique CDR3β amino acid sequences (a.k.a. 85% of the individuals were white, 52% were male, and 45% were female, with the remainder of unknown sex. Thus, we studied a cohort of 666 individuals, with annotated CMV serostatus, ethnicity, age, sex, high-resolution HLA class I and class II genotypes, and bulk TCRβ sequencing from PBMCs (Additional file 1: Table S1). However, it remains unknown to what extent HLA polymorphism affects TCR repertoire diversity in humans. These data, together with structural studies of the TCR-MHC interface, have provided key insights into how the TCR binds MHC and peptide. MHC restriction is the cornerstone of T cell recognition, and prior reports have assessed the effect of the presence of specific MHC alleles on TCR V gene usage and repertoire sharing. 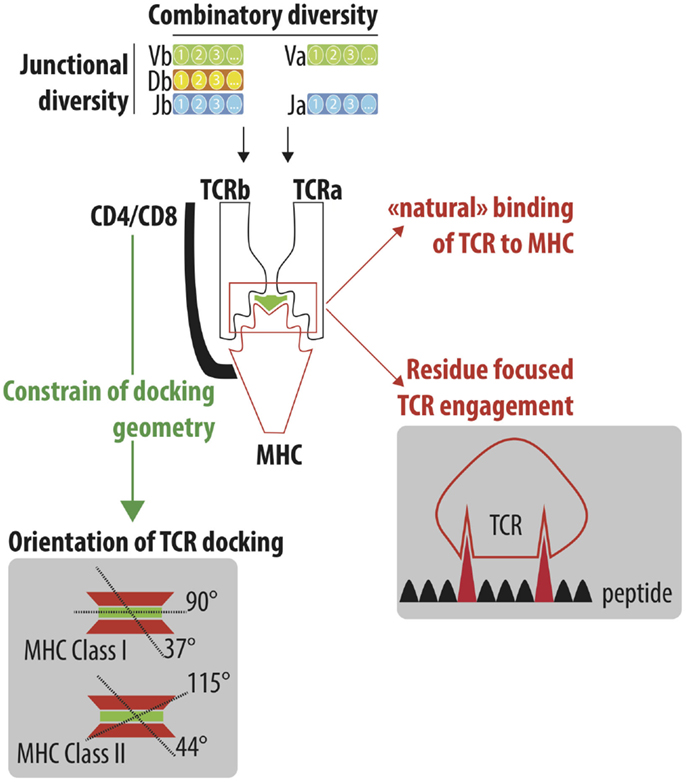
Accordingly, considerable effort has been devoted to understanding how MHC genetic variation impacts the TCR repertoire. TCR diversity can determine how efficiently one rejects pathogens such as viruses, and potentially cancer cells. 
Those TCRs that survive thymic selection are responsible for mounting productive immune responses through continuous interaction with self and foreign peptides bound to MHC molecules. TCRs that fail to bind to peptide-MHC complexes and those that bind too strongly are eliminated. Whether a particular TCR joins the periphery depends on its behavior during thymic selection, in which TCRs interact with both self peptide and MHC. The CDR3 regions of the TCR are primarily responsible for interacting with the peptide presented by MHC, with the potential diversity of CDR3β exceeding that of CDR3α. In the thymus, VDJ recombination facilitates random rearrangement of the complementary determining region 3 (CDR3) within the TCR α and β loci, followed by random nucleotide insertion and deletions at junction sites. This diversity, estimated to exceed 10 6 sequences in humans, is shaped by stochastic and genetic effects in conjunction with continuous immunological challenges throughout life. The large sequence diversity of the TCR repertoire is a hallmark of the adaptive immune system, and varies markedly across individuals. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
